Export Data¶
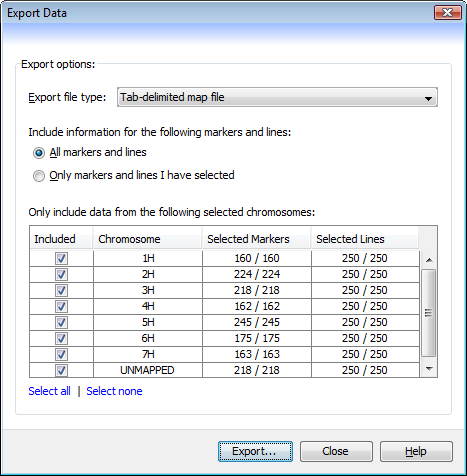
The Export Data dialog (Visualization->Export view as text) can be used to save the state of the current data set (maps, markers, and genotypes) as plain text files. The dialog will create either a .map file (for map data) or a .dat file (for genotype data).
The first step in outputting data is to decide what type of file to create.
Tab-delimited map file- selecting this option creates a .map file, that will contain a tab-delimited list of markers from the dataset. The file contains one marker per line, with each line containing three columns: the marker name, the name of the chromosome it is on, and its position (in cM) within that chromosome.Tab-delimited genotype file- selecting this option creates a .dat genotype file, that will contain a tab-delimited list of lines from the dataset. A header row references the markers, then each subsequent row contains the name of a line, followed by all the allele scores for that line against each marker.
The second and third steps allow you to filter the exported data, to include either all the information within the current data set, or just a subset of it.
All markers and lines- if selected, data on all markers and all lines will be exported.Only markers and lines I have selected- selecting this option ensures that only data on currently selected markers and lines will be exported. Any unselected markers or lines will be excluded from the export.Only include data from the following selected chromosomes- use the table to select which chromosomes should be included in the final output, with only those chromosomes that have theirIncludedfield ticked being used. Quickly include or exclude all the chromosomes by using theSelect allandSelect nonebuttons.
Mixing and matching these settings allows you to create files that include either all markers and lines (from either all or just some chromosomes), or just some markers and lines (again, from either all or just some chromosomes).
The exported files can either be re-imported into Flapjack, or can be opened and viewed in an external text editor or spreadsheet.